This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is gene ontology?
Gene Ontology is a bioinformatic resource that focuses on expanding the knowledge of genes, as well as greatly increasing efficiency of retrieval of such information.[1] The GO consortium is both human and machine readable, and thus allows for quick information retrieval as well as complex calculations. Gene ontology (GO) uses three domains to structure their resource and give an overall view of the gene: molecular function, cellular component, and biological process. [1]
Molecular Function
The functions and activities that the gene performs on a molecular level. Some examples are transporter activity, catalytic activity, or protein kinase activity. This does not specify which organelles do what or where this occurs.[1]
Cellular Component
The location where the gene product performs its activity or function. For example, gene products that function in the mitochondria would have mitochondria as their cellular component. [1]
Biological Processes
The larger processes that the gene product has an involvement in. For example, the gene that functions in the mitochondria may have involvement in metabolism. [1]
Molecular Function
The functions and activities that the gene performs on a molecular level. Some examples are transporter activity, catalytic activity, or protein kinase activity. This does not specify which organelles do what or where this occurs.[1]
Cellular Component
The location where the gene product performs its activity or function. For example, gene products that function in the mitochondria would have mitochondria as their cellular component. [1]
Biological Processes
The larger processes that the gene product has an involvement in. For example, the gene that functions in the mitochondria may have involvement in metabolism. [1]
TMC6 Gene Ontology
Cellular Component
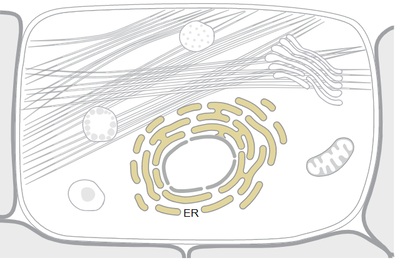
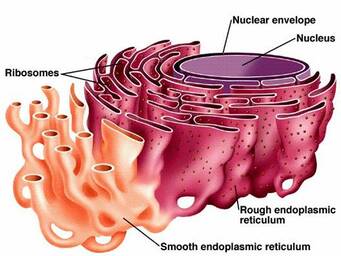
TMC6 is known to localize in the membrane of the Endoplasmic Reticulum (ER). [2] The ER is an extensive network of membrane tubules, vesicles, and flattened cisternae found throughout the cell. [3] The ER is made up of both the smooth and rough ER. The rough ER is lined with ribosomes, and this is the site of protein synthesis. The smooth ER lacks the ribosomes necessary to aide in protein synthesis, but it functions in lipid synthesis. It makes sense that loss of innate immunity to viral-protein synthesis would be located on the ER membrane.
Molecular Function
|
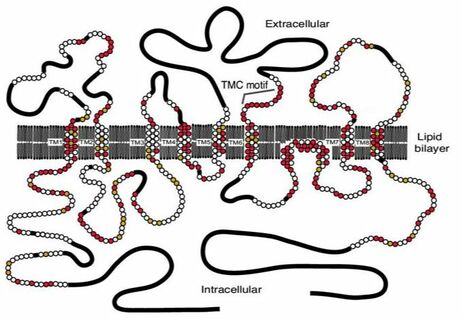
The full extent of how TMC6 functions in the cell is unknown; however, certain functions are well understood. It is known that TMC6 acts as an ion channel and has mechanosensitive ion channel activity. [2] It is hypothesized that TMC6 is involved with both Zinc-transport and intracellular chloride regulation. [4,5,6] It interacts with ZnT-1 to regulate zinc-transport and with calcium and integrin binding protein (CIB1) to regulate intracellular chloride concentrations. |
Biological Processes
TMC6 is known to be involved in neutrophil degranulation. [2] Neutrophil degranulation is the "regulated exocytosis of secretory granules containing preformed mediators such as lipases, proteases, and protein mediators". [7] This usually has a profound effect on the immune system, as many of the secretory vesicles contain cytotoxic molecules. Along with this, it is well understood that TMC6 is extensively involved in the innate cellular immune response to beta-HPV infection; however, it is unclear exactly how this works.
Conclusion
Gene ontology information is important to quickly and efficiently gather the necessary information regarding gene products. This is imperative to the study of TMC6 as it is a relatively unstudied gene. As new information updates GO, it allows scientists to always work with the most updated information on a gene's functions. Regarding TMC6 it gives an overall picture and idea of how it functions in the cell. This also elucidates possible ideas of how mutations in TMC6 can directly cause EV.
Sources:
1. http://geneontology.org/docs/ontology-documentation/
2. https://www.uniprot.org/uniprot/Q7Z403
3. https://www.uniprot.org/locations/SL-0097
4. Pryzbyszewska, J. (2016, December 21). Re-evaluation of epidermodysplasia verruciformis: Reconciling more than 90 years of debate. Retrieved February 8, 2019, from https://www-sciencedirect-com.ezproxy.library.wisc.edu/science/article/pii/S0190962216312968
5. Kalińska-Bienias, A., Kowalewski, C., & Majewski, S. (2016). The EVER genes - the genetic etiology of carcinogenesis in epidermodysplasia verruciformis and a possible role in non-epidermodysplasia verruciformis patients. Postepy dermatologii i alergologii, 33(2), 75-80.
6. de Jong, S. J., Crequer, A., Matos, I., Hum, D., Gunasekharan, V., Lorenzo, L., Jabot-Hanin, F., Imahorn, E., Arias, A. A., Vahidnezhad, H., Youssefian, L., Markle, J. G., and 25 others. The human CIB1-EVER1-EVER2 complex governs keratinocyte-intrinsic immunity to beta-papillomaviruses. J. Exp. Med. 215: 2289-2310, 2018.
7. https://www.ebi.ac.uk/QuickGO/term/GO:0043312
1. http://geneontology.org/docs/ontology-documentation/
2. https://www.uniprot.org/uniprot/Q7Z403
3. https://www.uniprot.org/locations/SL-0097
4. Pryzbyszewska, J. (2016, December 21). Re-evaluation of epidermodysplasia verruciformis: Reconciling more than 90 years of debate. Retrieved February 8, 2019, from https://www-sciencedirect-com.ezproxy.library.wisc.edu/science/article/pii/S0190962216312968
5. Kalińska-Bienias, A., Kowalewski, C., & Majewski, S. (2016). The EVER genes - the genetic etiology of carcinogenesis in epidermodysplasia verruciformis and a possible role in non-epidermodysplasia verruciformis patients. Postepy dermatologii i alergologii, 33(2), 75-80.
6. de Jong, S. J., Crequer, A., Matos, I., Hum, D., Gunasekharan, V., Lorenzo, L., Jabot-Hanin, F., Imahorn, E., Arias, A. A., Vahidnezhad, H., Youssefian, L., Markle, J. G., and 25 others. The human CIB1-EVER1-EVER2 complex governs keratinocyte-intrinsic immunity to beta-papillomaviruses. J. Exp. Med. 215: 2289-2310, 2018.
7. https://www.ebi.ac.uk/QuickGO/term/GO:0043312